Diffusion Autoencoders for Prediction of Sudden Cardiac Death
Using Representations from Diffusion Autoencoders for Prediction of Sudden Cardiac Death
Overview
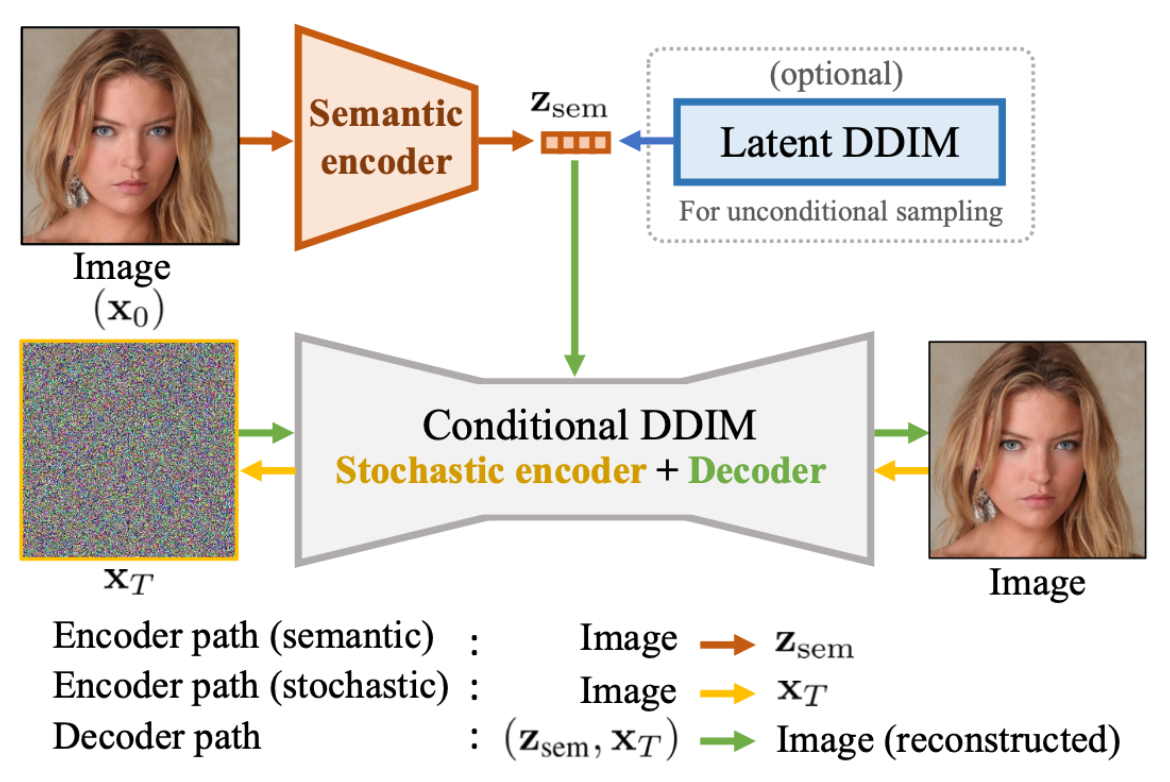
I adapted diffusion autoencoders, originally designed for facial image datasets, to work with medical imaging data. The model combined a semantic encoder (UNet architecture) for capturing high-level features with a diffusion model component (Denoising Diffusion Implicit Model - DDIM) for handling fine-grained stochastic details. The approach was tested on two datasets: the publicly available Emidec dataset (100 samples) and the DEEP-RISK dataset from Amsterdam UMC (924 samples).
Technical Approach
The semantic encoder used a modified UNet architecture with attention mechanisms and time conditioning. The stochastic component utilized DDIM, which offers deterministic reconstruction and requires fewer inference steps than standard diffusion models. The key innovation was conditioning the diffusion decoder on semantic encodings through Adaptive Group Normalization (AdaGN), allowing high-level semantic information to guide the denoising process.
For the medical imaging application, I modified the model to handle single-channel grayscale images (128×128 pixels) instead of RGB images, and implemented preprocessing to crop images around the myocardium region using provided contours. Training used the simplified DDPM loss with 1000 diffusion steps during training and 20 steps during evaluation.
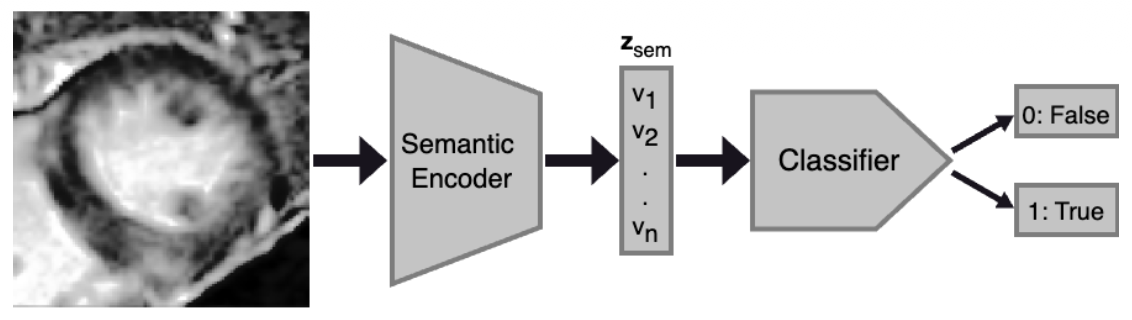
Data and Evaluation
Reconstruction performance was evaluated using Structural Similarity Index (SSIM) and Mean Squared Error (MSE). Classification performance used AUROC, F1-score, precision, and recall metrics, with a multilayer perceptron classifier trained on the semantic encoder's latent representations.
Key Outcomes
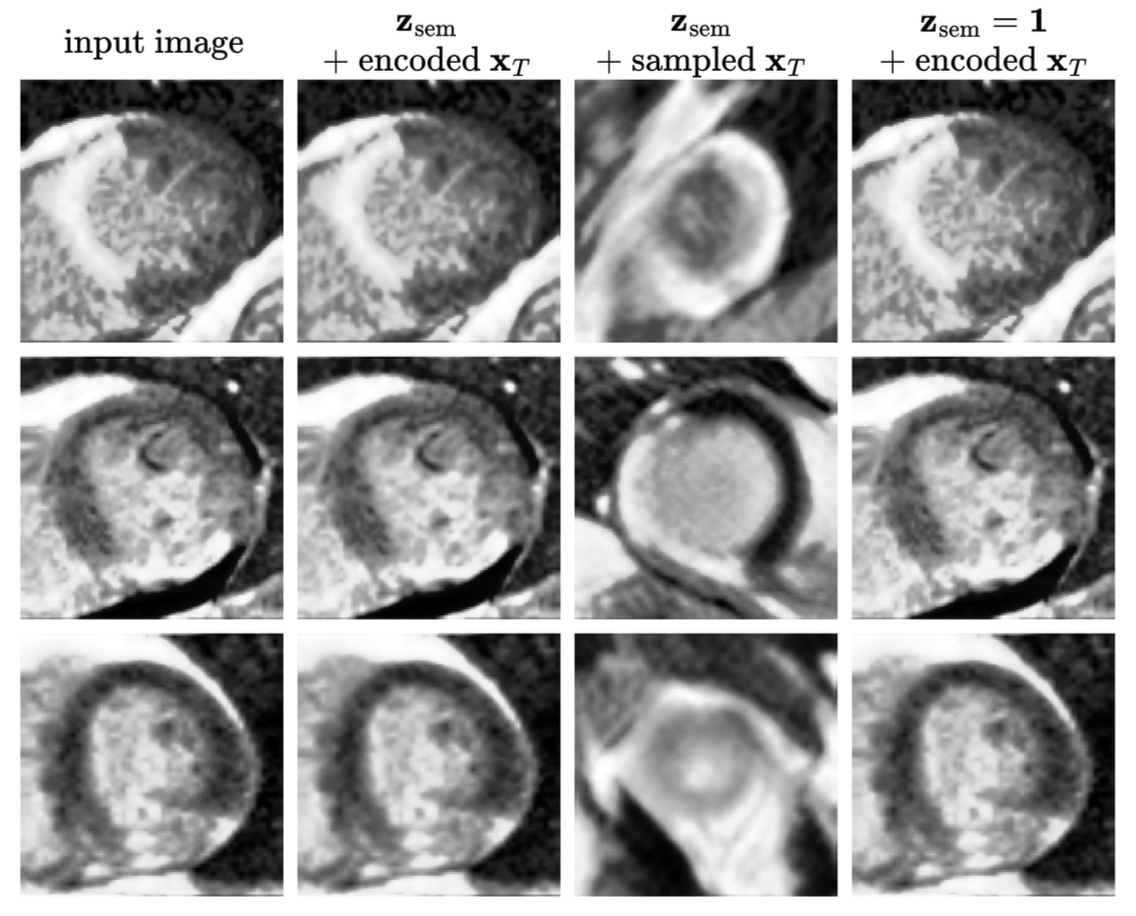
- When using both semantic and stochastic encodings, reconstruction achieved SSIM of 0.66 (Emidec) and 0.65 (DEEP-RISK), outperforming VAE baselines
- Using only semantic encodings with sampled stochastic representations resulted in dramatically degraded reconstruction: SSIM of 0.20 (Emidec) and 0.17 (DEEP-RISK), 91-202% worse MSE than VAE models
- Classification performance was substantially inferior to VAE-based approaches across all metrics, with F1-scores of 0.78 (Emidec) and 0.47 (DEEP-RISK) for single-slice classification
- Extensive training (5-10x longer) improved reconstruction metrics but classification remained significantly below VAE performance
Most Interesting Findings
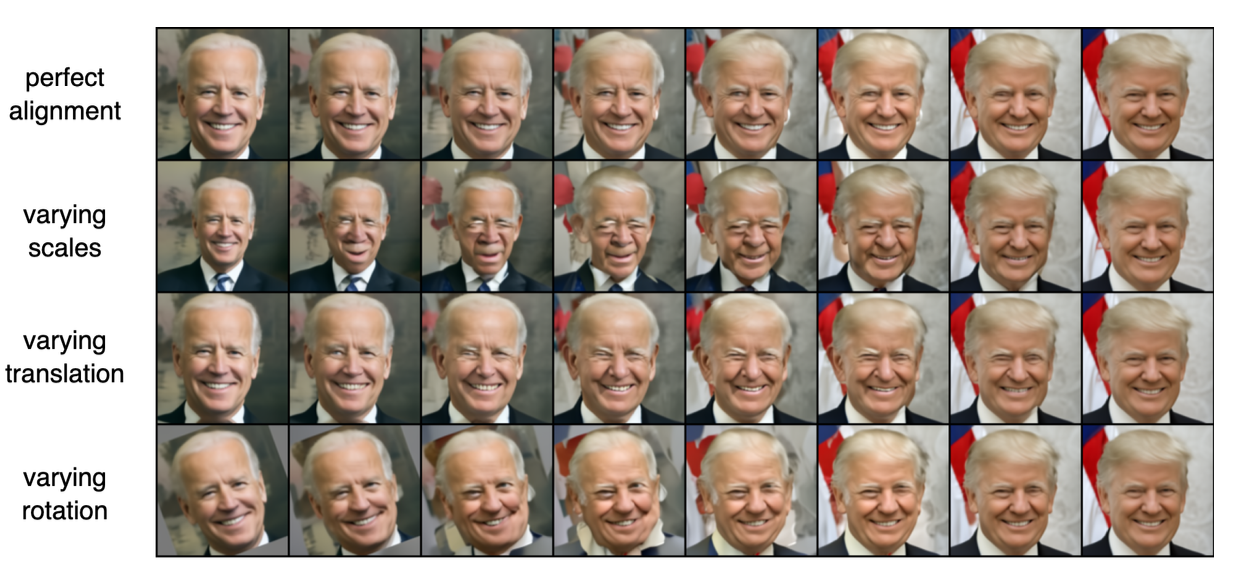
The most significant discovery was that the semantic encoder failed to capture meaningful global features of the cardiac images, contrary to the model's performance on facial datasets. Visual inspection revealed that reconstructions relied almost entirely on the stochastic encoding, with the semantic encoding providing negligible benefit—a fundamental departure from the model's intended behavior. This failure was attributed to the diffusion autoencoder's high sensitivity to geometric transformations and inconsistent feature locations across the medical imaging datasets. The research revealed a critical limitation: the significantly smaller dataset size (800-7,392 images) compared to the original diffusion autoencoder training data (60,000 images) likely contributed to the suboptimal performance. While diffusion autoencoders excel at high-quality reconstruction when all information is available, the semantic encoder did not learn to encode diagnostically relevant features for cardiac pathology classification. An interesting technical finding was that scaling the semantic encodings in the conditioning process improved both reconstruction and classification performance, particularly for the Emidec dataset, achieving the highest AUROC scores in the final experiments. This suggested that architectural modifications to enhance the semantic encoder's influence could partially address the observed limitations.
Practical Implications
The project demonstrated important limitations of applying state-of-the-art generative models from computer vision to medical imaging domains. While the diffusion autoencoder maintained high reconstruction fidelity with complete information, it failed to provide the interpretable, semantically meaningful latent representations necessary for clinical decision-making. The research highlighted the need for improved preprocessing to ensure feature locality consistency and potential architectural refinements to enhance robustness against geometric variations in medical imaging. These findings have important implications for the medical AI community, emphasizing that models successful in natural image domains require substantial adaptation and validation before deployment in clinical settings where interpretability and reliability are paramount.
